First-Strand cDNA Synthesis – Notes and Outcome
Overview
We set up first-strand cDNA synthesis from total RNA extracted from zebrafish liver and larvae as part of a bulk 3′ RNA-seq workflow.
RNA used in this protocol was extracted using the Promega ReliaPrep Tissue MiniPrep System (Z6111).
Materials
Consumables
- RNase/DNase-free PCR tubes
- RNase/DNase-free 1.5 mL tubes
- RNase/DNase-free pipette tips
Reagents
- MMLV Reverse Transcriptase (2 mg/mL, in-house or equivalent)
- 5× First Strand Buffer (ThermoFisher, Cat# Y02321)
- DTT (0.1 M, included with buffer)
- Anchored Oligo(dT)20 (100 µM) (Jena, PM-305)
- dNTP mix (25 mM) (Jena, NU-1023)
- RNase inhibitor (Jena, PCR-392)
- DNA Purification Magnetic Beads (ABM, G951 or SPRI beads or equivalent)
- EtOH
- EDTA (0.5 M, pH 8.0)
Buffer composition (5× First Strand Buffer):
- 250 mM Tris-HCl (pH 8.3)
- 375 mM KCl
- 15 mM MgCl₂
General Notes
- Work under RNase-free conditions
- RNA quality is critical (RIN > 8 recommended)
- Input: 1 µg total RNA (~100 ng/µL)
Procedure
1. Reaction Setup (Part 1)
In a PCR tube:
| Component | Volume |
|---|---|
| RNA (1 µg) + water | 10.7 µL |
| 5× First Strand Buffer | 4 µL |
| Oligo(dT)20 (100 µM) | 2 µL |
| dNTPs (25 mM) | 0.8 µL |
Thermocycler:
- 65°C – 5 min
- 4°C – hold
2. Reaction Setup (Part 2)
Add to the same tube:
| Component | Volume |
|---|---|
| Previous mix | 17.5 µL |
| RNase inhibitor | 0.5 µL |
| DTT (0.1 M) | 1 µL |
| MMLV RT | 1 µL |
Thermocycler:
- 55°C – 60 min
- 70°C – 10 min (inactivation) [Do not perform if the goal is to generate RNA-Seq libraries]
- 4°C – hold
Optional: EDTA Stabilization
To protect cDNA:
- Add 0.1 mM EDTA final concentration
Preparation:
- Dilute 0.5 M EDTA → 5 mM (1:100)
- Add 0.4 µL of 5 mM EDTA to 20 µL reaction
Note: Higher EDTA can inhibit downstream reactions due to Mg²⁺ chelation.
Storage
- Short-term: 4°C (≤24 hours)
- Long-term: −20°C
- Avoid repeated freeze–thaw cycles
Bead-Based Purification
Before downstream applications, perform bead-based cleanup of cDNA:
- Bring cDNA to room temperature.
- Add 1.8× volume of magnetic beads (e.g., 36 µL beads to 20 µL cDNA).
- Mix thoroughly and incubate for 5–10 minutes at room temperature.
- Place on magnetic stand and allow beads to separate (~5 minutes).
- Remove supernatant carefully without disturbing beads.
- Wash beads twice with 80% ethanol (200 µL each), without resuspending.
- Air-dry beads for 2–5 minutes (do not overdry).
- Elute cDNA in 10–20 µL nuclease-free water.
- Incubate 2 minutes, place on magnet, and collect eluate.
- Measure concentration using Nanodrop. Expected range: 10–50 ng/µL.
Purified first-strand cDNA can be used directly for PCR or library preparation.
Validation by PCR
Dilute cDNA in nuclease-free water such that the final concentration is 1–5 ng/µL (typically ~1:20 dilution, adjust based on measured concentration).
Set up a standard endpoint PCR:
| Component | Volume |
|---|---|
| cDNA (1:20 diluted) | 1 µL |
| Forward primer (10 µM) | 0.4 µL |
| Reverse primer (10 µM) | 0.4 µL |
| dNTPs (10 mM) | 0.4 µL |
| 10× buffer | 2 µL |
| Polymerase | 0.2–0.5 µL |
| Water | to 20 µL |
Run on agarose gel to confirm amplification.
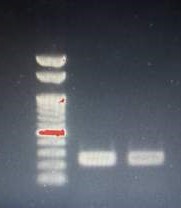
W1: Ladder
W2: cDNA from Adult Liver
W3: cDNA from 5 dpf Larvae
Notes
- RNase inhibitor can be increased (up to 1 µL) if RNA input is low
- Can be omitted if RNase-free conditions are well controlled
- In-house MMLV (~2 mg/mL) corresponds roughly to 200–400 U/µL
- Typical usage: 100–200 U per reaction (0.5–1 µL)
Summary Workflow
Total RNA → Denaturation → Reverse Transcription → Bead Cleanup → cDNA → PCR Validation
Contributors
- Ishika Jain
- Kaushani Ghosh
- Namra Ali
Copyright & License
© 2026 Singh Lab and Omics Facility, Shiv Nadar University.
This document is distributed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
This license permits use, distribution, adaptation, and reproduction in any medium or format, provided that appropriate credit is given to the authors and the Singh Lab and Omics Facility, Shiv Nadar University, and any modifications are clearly indicated.
License details: https://creativecommons.org/licenses/by/4.0/